Spatial Defects in 5896 HG-U133A GeneChips
W. B. Langdon
Mathematical and Biological Sciences
Essex University
Presented at
CAMDA-2007
Introduction
- Examples of spatial defects
- NCBI GEO and
E-TABM-185 human GeneChips
- Non-parametric detection of spatial flaws
- Error rate for each GeneChip type
- Where next
Example Laser output
Next

Example GeneChip Flaw visible in Laser Scanner output
back
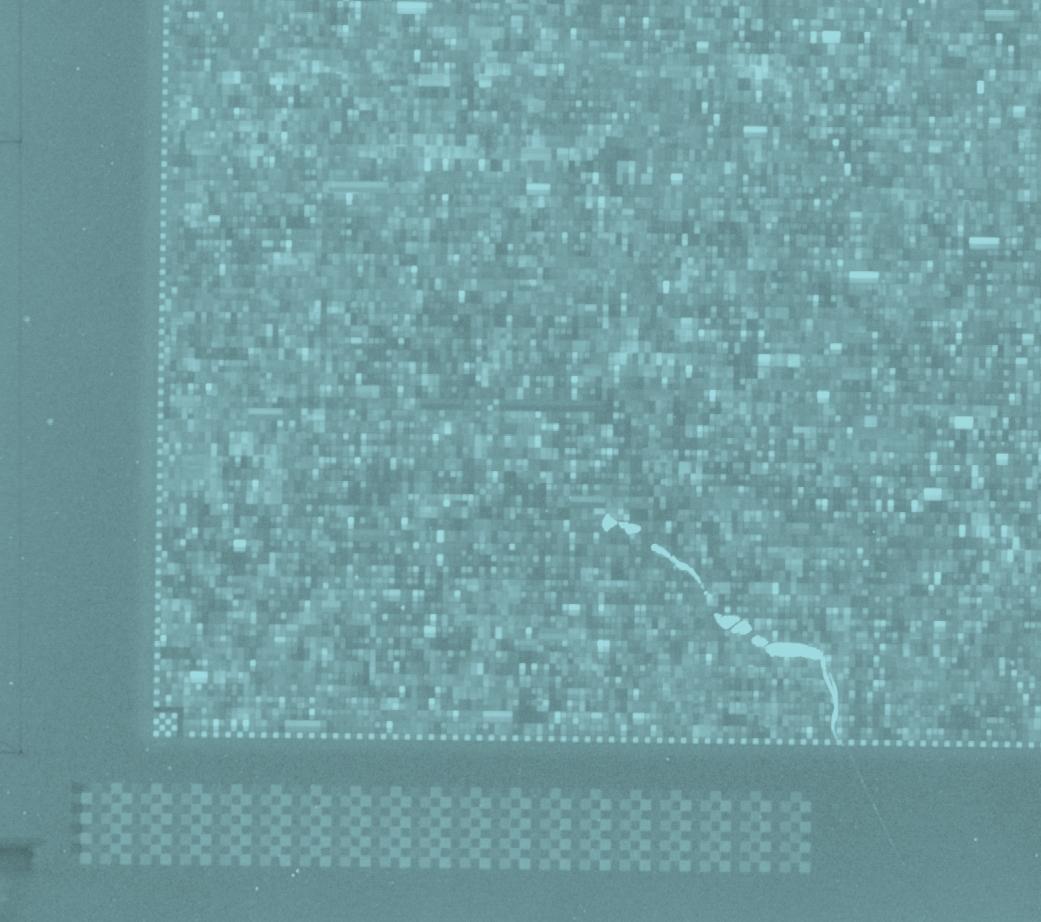

Example flaw in CEL file

Example large area flaw in CEL file
First CEL file in GSM2394

Quantile normalised log scaled
Red suspiciously bright
Detecting Spatial Flaws

- Are group of probes all above average?
- All below average?
- 3 by 3 checker board.
- Count number of coloured squares which have same sign as centre.
- For statistical independence ignore vertical and horizontal neighbours.
- Chequer board used to give statistical independence by ensuring don't use both perfect match (PM) and mismatch (MM) probes.
- 50% above average.
- p(all above average) = 1/32
- Millions of probes, 1/32 too often by chance.
Hierarchical test. See if adjacent 3x3 are also all high (or all low)
Detecting Spatial Flaws

- 9 3x3 checker boards.
-
If 3 or more 3x3 neighbours are above average,
Flag central 3x3 as being suspicious.
- P(≥3) <0.01%
Mean 5896 E-TABM-185
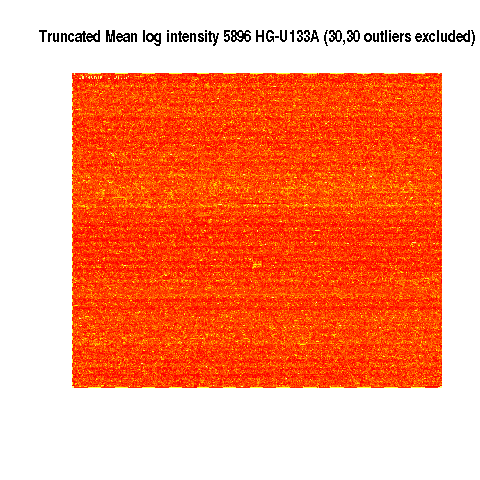 Yellow is highest
Yellow is highest
Example E-TABM-185 HG-U133A
Choose as example HG-U133A with average number of flaws.

Quantile Normalised
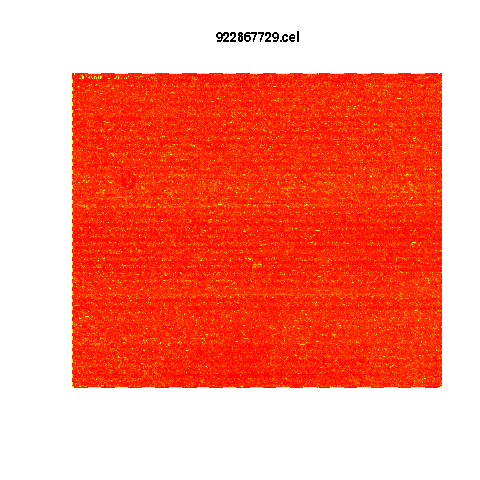 Yellow is highest
Yellow is highest
Example HC-U133A v average
 Yellow is highest
Yellow is highest
Flaws in example E-TABM-185
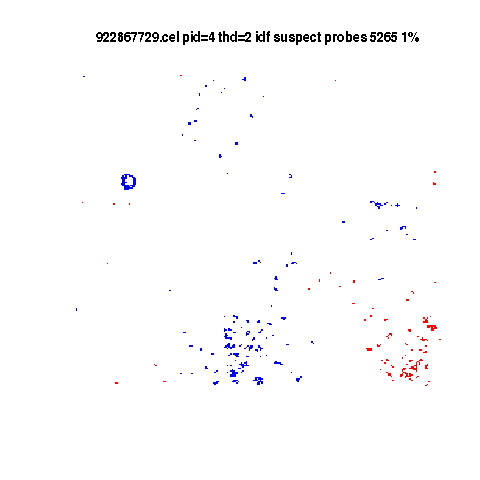 Blue below average
Blue below average
Red above average.
Controls suppressed
Defects Matter
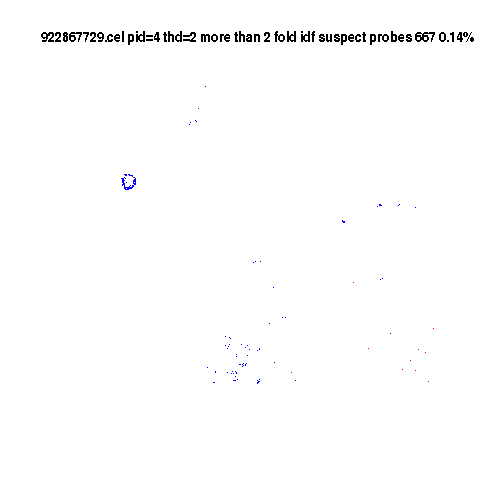 Blue less than half expected. Min=1/6
Blue less than half expected. Min=1/6
Red more than 2 fold more than expected. Max 17 times average.
Controls suppressed
Distribution of suspect probes in 5896 E-TABM-185
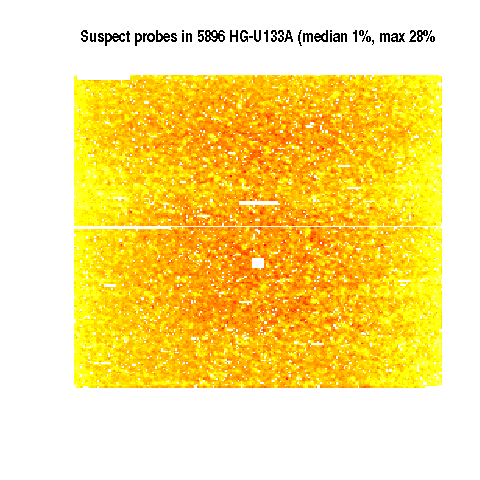 Yellow is highest
Yellow is highest
Controls suppressed
GEO/E-TABM-185 defect rate
Summary
- More than 15000 GeneChips analysed.
- All published human chips have errors.
- Uneven error rate. Some probes 28%
- Error rate depends on chip type but has fallen.
- Method for detecting spatial defects. Applied to chips one at a time.
- Technique is non-parametric, does not assume Gaussian or other distributions, is statistically sound, fast and in R.
(6000 HG-U133A processed in 5hours 18minutes on desk top Linux computer).
END
Mean 5896 E-TABM-185
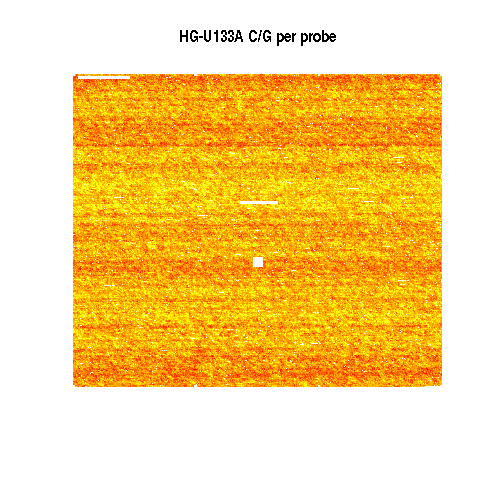 Yellow is highest
Yellow is highest
W.Langdon
17 Dec 2007














